
A powerful and intuitive standalone interface for molecular docking, HTVS, pharmacophore modeling, protein–protein docking, membrane system workflows, molecular dynamics workflow management, and trajectory analysis within a streamlined computational drug discovery environment.
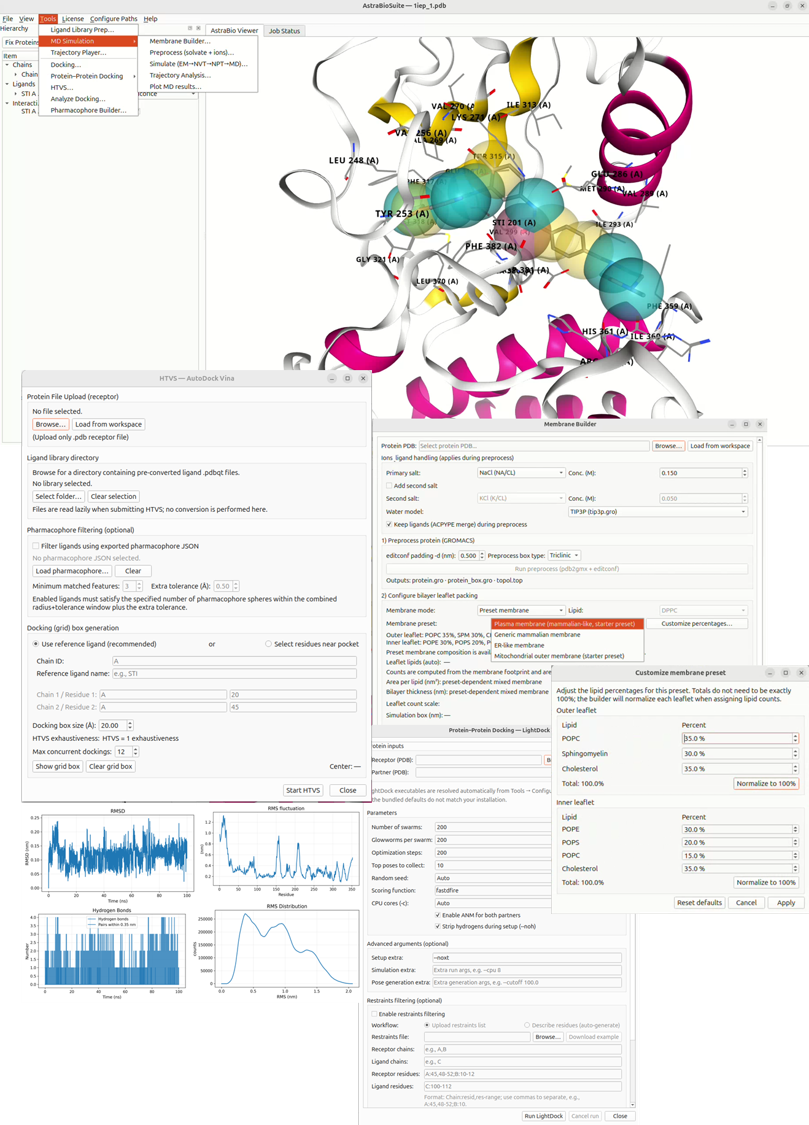
AstraBioSuite™ brings integrated workflow interfaces (UI) for molecular docking, ligand-based screening, pharmacophore modeling, protein–protein docking, membrane system preparation, GROMACS-based molecular dynamics workflows, and trajectory analysis into one streamlined computational drug discovery environment.
Modern computational drug discovery often requires several disconnected tools, manual file conversions, command-line setup, and complex troubleshooting. AstraBioSuite™ is designed to reduce this fragmentation by offering an intuitive standalone interface for structure-based, ligand-based, simulation-based, and membrane-aware modeling workflows.
The UI platform supports molecular docking, high-throughput virtual screening, pharmacophore model generation, pharmacophore-based HTVS, protein–protein docking, GROMACS-based molecular dynamics simulations, preset membrane system workflows, and post-simulation trajectory analysis.
A key differentiator of AstraBioSuite™ is its integrated preset membrane system workflow, including mammalian-like plasma membrane, ER-like membrane, and mitochondrial outer membrane environments. These preset systems help researchers build biologically relevant membrane models for GPCRs, transporters, receptors, ion channels, viral proteins, and mitochondrial targets.
From ligand screening to membrane simulations, AstraBioSuite™ helps researchers move from molecular model to simulation-ready system with fewer fragmented steps.
Built for researchers who need integrated, biologically relevant, and user-friendly computational modeling workflows.
Perform structure-based docking workflows with guided receptor preparation, ligand handling, grid setup, docking execution, and binding pose interpretation.
Screen compound libraries through streamlined HTVS workflows for hit identification, compound ranking, and downstream lead prioritization.
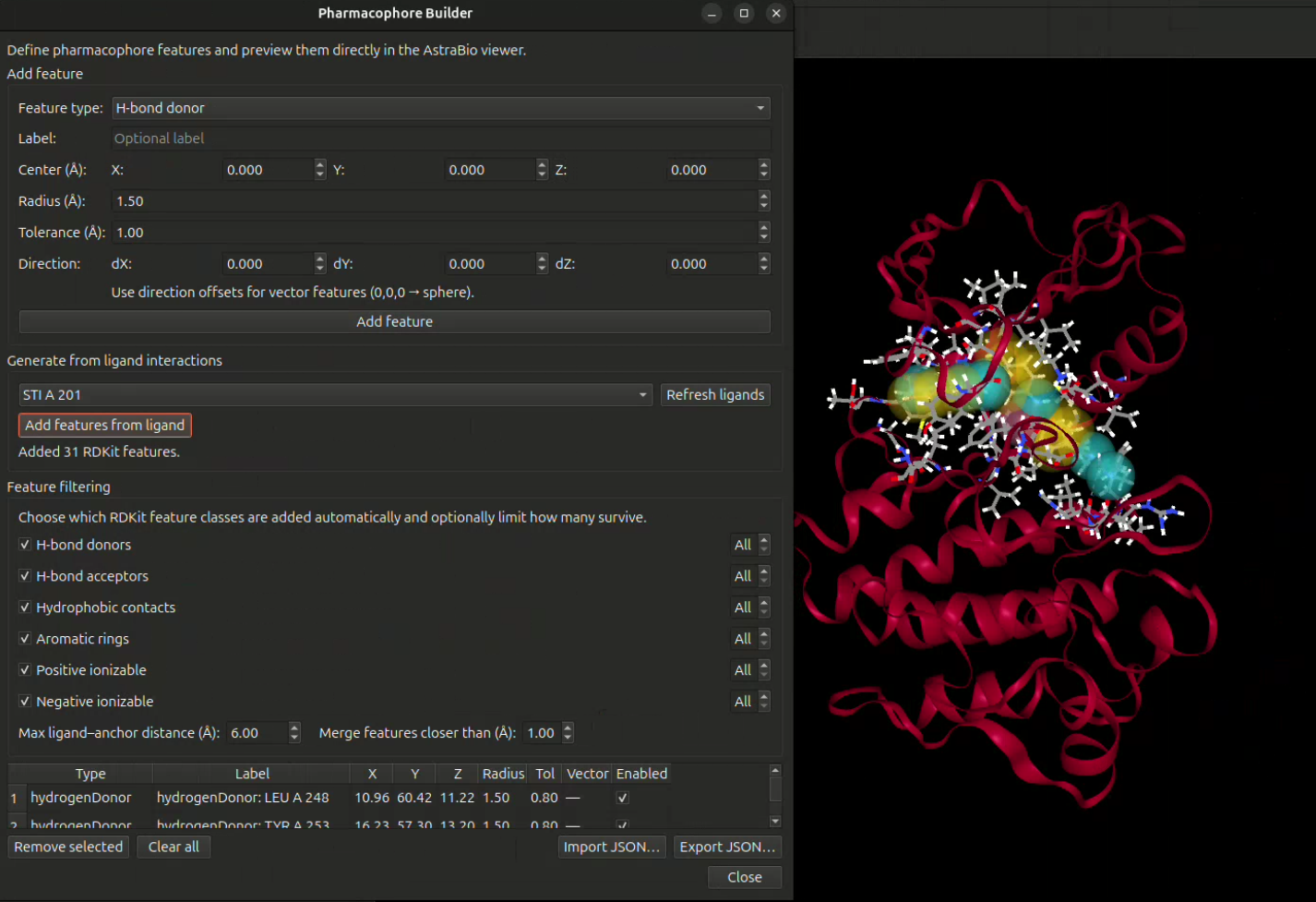
Generate pharmacophore models from ligand features or protein–ligand interaction patterns, including key hydrogen bond donors, acceptors, hydrophobic regions, aromatic features, and charged interaction points.
Use pharmacophore models to screen compound libraries and prioritize molecules that match essential interaction features before docking or simulation-based validation.
Explore protein–protein interaction models with workflows designed for interface prediction, complex generation, interaction analysis, and refinement-ready structural outputs.
Build biologically relevant membrane simulation systems using integrated presets for mammalian-like plasma membrane, ER-like membrane, and mitochondrial outer membrane environments.
Prepare and launch molecular dynamics workflows through a user-friendly interface designed to support GROMACS-based simulations for proteins, protein–ligand complexes, and membrane-associated targets.
Analyze simulation trajectories using essential post-MD metrics including RMSD, RMSF, hydrogen bonds, SASA, and structural stability indicators.
View molecular models, docking poses, protein–ligand complexes, and simulation outputs within a researcher-friendly interface designed for easier interpretation.
AstraBioSuite™ expands beyond docking by enabling pharmacophore-guided compound discovery and feature-based virtual screening workflows.
The pharmacophore module helps identify essential molecular interaction features such as hydrogen bond donors, hydrogen bond acceptors, hydrophobic centers, aromatic features, positive ionizable groups, negative ionizable groups, and spatial feature arrangements important for target binding.
Researchers can use pharmacophore models to screen compound libraries and prioritize molecules that satisfy key interaction requirements before advancing to molecular docking, molecular dynamics, or membrane-based simulation workflows.
Pharmacophore-based HTVS provides an additional ligand-based discovery strategy, helping users identify structurally diverse compounds that preserve important binding features even when their chemical scaffolds differ.
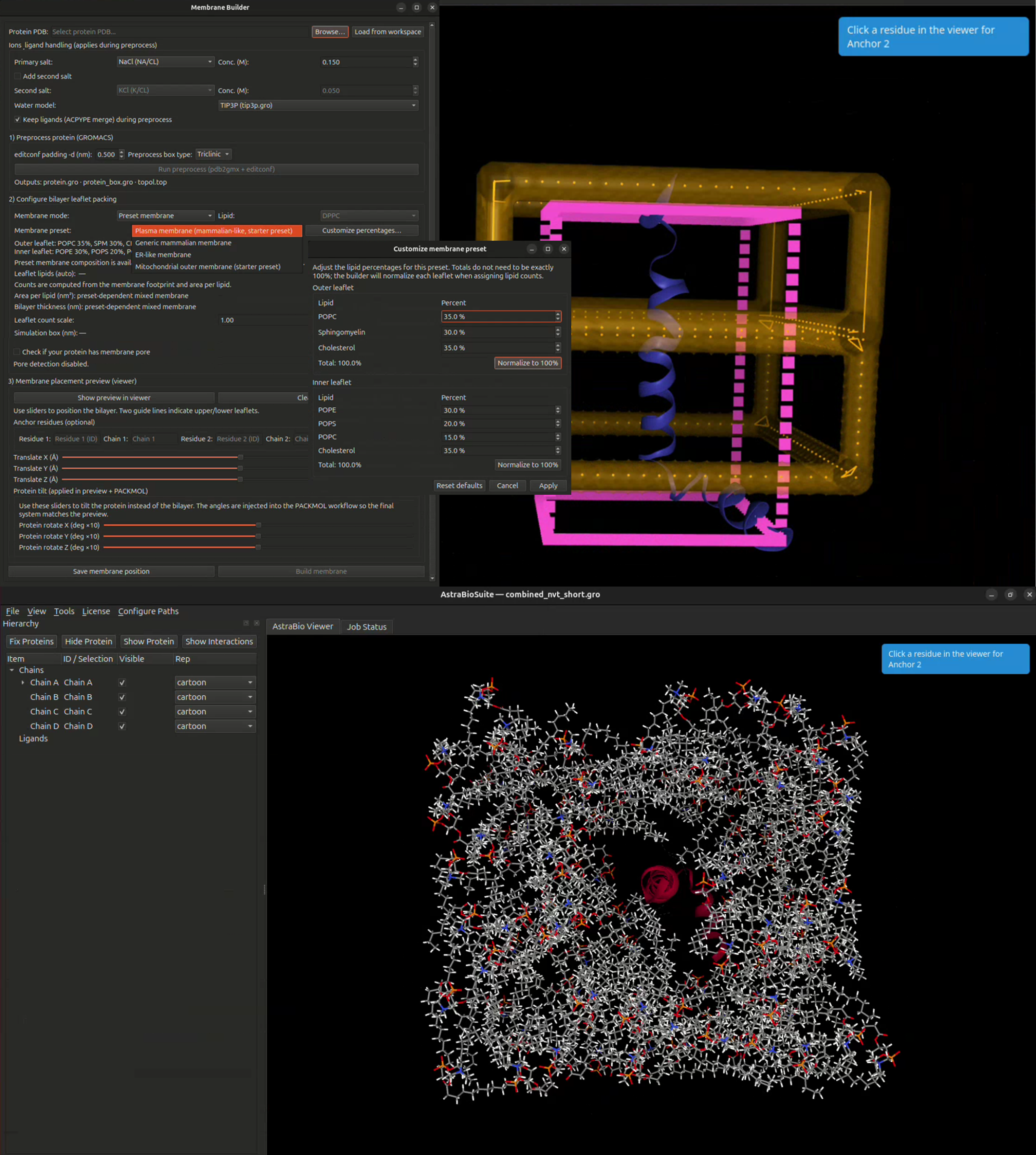
Membrane preparation is often one of the most technically challenging steps in molecular dynamics. AstraBioSuite™ simplifies this process with integrated preset membrane system workflows.
AstraBioSuite™ includes preset membrane environments designed to support biologically relevant molecular simulations for membrane proteins and membrane-associated therapeutic targets.
Designed for studies involving cell-surface receptors, GPCRs, ion channels, transporters, viral entry proteins, and ligand–receptor interactions at the plasma membrane.
Useful for modeling proteins associated with the endoplasmic reticulum environment, including ER-resident proteins, membrane-associated folding pathways, lipid interactions, and organelle-specific protein behavior.
Supports simulation setup for mitochondrial outer membrane-associated proteins, apoptosis-related targets, metabolite transport systems, and mitochondrial signaling proteins.
These preset membrane workflows reduce manual setup time and help researchers move more efficiently from protein structure to simulation-ready membrane systems.
AstraBioSuite™ supports computational drug discovery from early-stage screening to simulation-based structural validation. The platform is designed to help researchers organize and execute molecular modeling tasks in a more streamlined and reproducible way.
Prepare small-molecule libraries for docking, HTVS, and pharmacophore-based screening workflows with simplified ligand handling and structure preparation support.
Run molecular docking and high-throughput virtual screening workflows for target-based hit discovery, compound prioritization, binding pose analysis, and downstream lead selection.
Use pharmacophore modeling and pharmacophore-based HTVS to identify compounds that preserve key molecular interaction features, enabling ligand-based screening in addition to structure-based docking.
Support protein–protein docking workflows for studying macromolecular interactions, interface regions, and complex formation.
Build membrane-associated simulation systems using preset mammalian-like plasma membrane, ER-like membrane, and mitochondrial outer membrane workflows for biologically relevant molecular dynamics studies.
Use molecular dynamics workflow management and trajectory analysis tools through a streamlined interface designed to support GROMACS-compatible simulation workflows for evaluating structural stability, conformational behavior, hydrogen bonding, surface exposure, and protein–ligand interaction patterns.
SiBioLEAD LLC provides an optional installation utility script for simplifying the setup of commonly used open-source bioinformatics and molecular modeling tools.
SiBioLEAD LLC does not redistribute any third-party software packages. Instead, the platform provides an integrated workflow environment designed to interface with selected open-source computational biology and molecular modeling tools.
To simplify software environment setup for researchers, SiBioLEAD LLC provides a freely downloadable installation utility script (install_bioinformatics-tools.sh) designed to help automate the setup of commonly used Linux-based open-source bioinformatics and molecular modeling command-line tools.
These third-party tools are independently developed open-source software packages that operate independently of AstraBioSuite™ and remain fully functional as standalone applications within their respective computational workflows. SiBioLEAD LLC does not claim ownership of any third-party open-source software referenced, installed, or accessed through this installation utility script .
Users are responsible for reviewing and complying with the individual license terms, usage policies, and citation requirements associated with each third-party software package.
The freely provided installation utility script is intended solely to assist with software environment setup for commonly used open-source computational biology tools and does not require the use, purchase, or adoption of AstraBioSuite™.